Machine Learning 101
The demand for data scientists is very clear in all industries, to include healthcare. While statistics has been the backbone of data analytics it is not the only approach to predictive modeling. Not only can a data scientist use the programming languages R and Python they can rely on machine learning (ML), traditionally the tool of computer scientists.The purpose of this blog is to point out that machine learning tools can now be in the domain of almost anyone. Users can choose from commercial products such as Microsoft SQL Server Analysis Services (SSAS) or IBM SPSS Modeler, but they can also opt to use free open-source applications. The following is a list of some of these open-source choices: RapidMiner, KNIME, Pentaho, Orange and WEKA. Many of these choices require dragging a widget (operator) onto a field and connecting the input with output, in order to perform a function. The exception to that is WEKA that was developed at the University of Waikato in New Zealand. Not only is their software widely used, it is complemented by a textbook, an online self paced course and multiple YouTube instructional videos.For those reasons I would like to focus on WEKA in this blog. The software is available for multiple operating systems and the installation is simple. The input must be either a .csv file or an .arff file. Under tools there is an .arff viewer that can view either file type and convert an .arff file to .csv, if needed. I commonly use a .csv file to run the data in both IBM Watson Analytics for the statistical approach and WEKA for the machine learning approach.Machine learning performs many functions, but the three most commonly used are classification, regression and clustering. Classification is used for predictions when the outcome is categorical data, such as lived or died (binary). Regression is used for scenarios where the outcome (or dependent variable) is continuous or numerical, such as healthcare cost in dollars. Clearly, one of the challenges here is the fact that computer scientists often use terms that are different from statisticians, but mean the same (column = features = attributes) (outcome = class = target = dependent variable). Once you can get by the terminology, the approach gets much simpler. WEKA comes with multiple sample data sets, the user can train on that are classic validated data sets. More data sets can be found on the UC Irvine Machine Learning Repository. Below is a screenshot of WEKA explorer (figure 1)Figure 1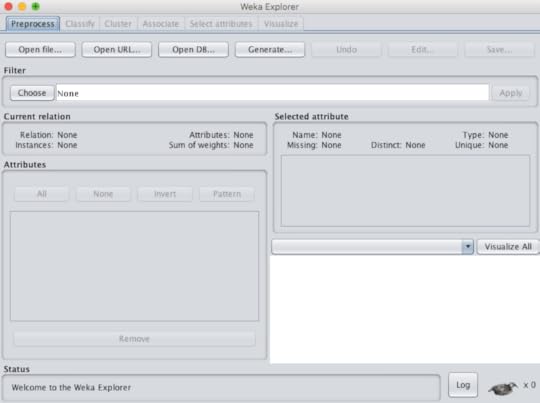 If the user wants to try out classification, they must first upload the file (.csv or .arff) under the "preprocess" tab. There they can perform "exploratory data analysis or EDA" by viewing the rows (instances) and features (attributes). This will provide an idea if the data is normally distributed or skewed. There is also a filter option, where data can be "normalized" or "standardized" and then analyzed. This can be important if your attributes are on very different scales. (figure 2)Figure 2
If the user wants to try out classification, they must first upload the file (.csv or .arff) under the "preprocess" tab. There they can perform "exploratory data analysis or EDA" by viewing the rows (instances) and features (attributes). This will provide an idea if the data is normally distributed or skewed. There is also a filter option, where data can be "normalized" or "standardized" and then analyzed. This can be important if your attributes are on very different scales. (figure 2)Figure 2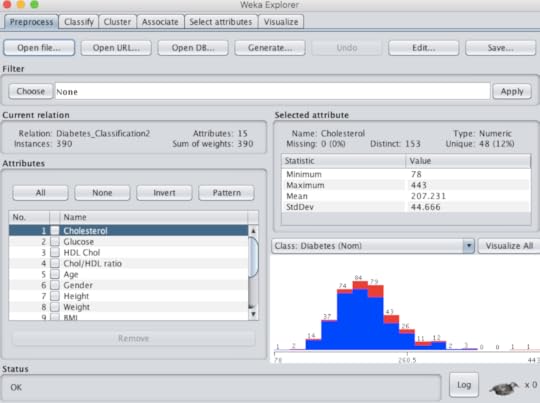 The "classify" tab is where the fun begins. Selecting the "Choose" option reveals an entire tree of algorithm choices. Those inappropriate for classification are greyed out. Unlike statistics, where normally one approach only is chosen, users routinely run several algorithms (like recipes) to see what approach or model provides the best results. For example, for most classification exercises I run a decision tree, Naive Bayes, logistic regression, K-nearest neighbor and support vector machine. Each run only takes one second or less. The results are very useful because it includes: classification accuracy, precision, recall (sensitivity), confusion matrix and area under the curve (AUC). Most healthcare workers are used to seeing results, in terms of sensitivity and specificity. If the results are similar they can be compared in another tab of WEKA called "experimenter" to determine if there is any statistical difference. Furthermore, there is another tab called "select attribute" where the user can determine which attributes really contribute to the outcome, as frequently only a few are really contribute to the outcome. Each algorithm can be further customized to see if the change affects the results. Below is the result of classification to determine what predicts type 2 diabetes in adults. We started with 14 predictors but ended with the best 4. (TP = true positive, FP = false positive) (table 1)Table 1
The "classify" tab is where the fun begins. Selecting the "Choose" option reveals an entire tree of algorithm choices. Those inappropriate for classification are greyed out. Unlike statistics, where normally one approach only is chosen, users routinely run several algorithms (like recipes) to see what approach or model provides the best results. For example, for most classification exercises I run a decision tree, Naive Bayes, logistic regression, K-nearest neighbor and support vector machine. Each run only takes one second or less. The results are very useful because it includes: classification accuracy, precision, recall (sensitivity), confusion matrix and area under the curve (AUC). Most healthcare workers are used to seeing results, in terms of sensitivity and specificity. If the results are similar they can be compared in another tab of WEKA called "experimenter" to determine if there is any statistical difference. Furthermore, there is another tab called "select attribute" where the user can determine which attributes really contribute to the outcome, as frequently only a few are really contribute to the outcome. Each algorithm can be further customized to see if the change affects the results. Below is the result of classification to determine what predicts type 2 diabetes in adults. We started with 14 predictors but ended with the best 4. (TP = true positive, FP = false positive) (table 1)Table 1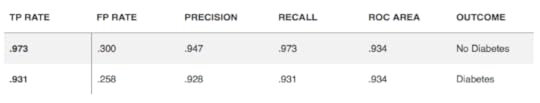 For regression, users select the "classify" tab and choose linear regression and for clustering they use the "cluster" tab. Clustering is unsupervised learning where you don't know the classes or attributes so you run clustering to see if there are groups hidden in the data that are worth further exploration. There is also a "select attributes" tab that determines which attributes are most important.I encourage healthcare workers in the data analytics realm to become familiar with WEKA or a similar program, as I believe they will find the programs useful and not that difficult to use.It is true that you must have some background information and most do not want to read an extensive textbook to gain this information. I highly recommend a 2016 ebook (248 page PDF) from Jason Brownlee that teaches basic machine learning and uses WEKA as the software application. It is written simply and clearly and doesn't involve coding or formulas! The cost is only $27 and is well worth it.
For regression, users select the "classify" tab and choose linear regression and for clustering they use the "cluster" tab. Clustering is unsupervised learning where you don't know the classes or attributes so you run clustering to see if there are groups hidden in the data that are worth further exploration. There is also a "select attributes" tab that determines which attributes are most important.I encourage healthcare workers in the data analytics realm to become familiar with WEKA or a similar program, as I believe they will find the programs useful and not that difficult to use.It is true that you must have some background information and most do not want to read an extensive textbook to gain this information. I highly recommend a 2016 ebook (248 page PDF) from Jason Brownlee that teaches basic machine learning and uses WEKA as the software application. It is written simply and clearly and doesn't involve coding or formulas! The cost is only $27 and is well worth it.
 If the user wants to try out classification, they must first upload the file (.csv or .arff) under the "preprocess" tab. There they can perform "exploratory data analysis or EDA" by viewing the rows (instances) and features (attributes). This will provide an idea if the data is normally distributed or skewed. There is also a filter option, where data can be "normalized" or "standardized" and then analyzed. This can be important if your attributes are on very different scales. (figure 2)Figure 2
If the user wants to try out classification, they must first upload the file (.csv or .arff) under the "preprocess" tab. There they can perform "exploratory data analysis or EDA" by viewing the rows (instances) and features (attributes). This will provide an idea if the data is normally distributed or skewed. There is also a filter option, where data can be "normalized" or "standardized" and then analyzed. This can be important if your attributes are on very different scales. (figure 2)Figure 2 The "classify" tab is where the fun begins. Selecting the "Choose" option reveals an entire tree of algorithm choices. Those inappropriate for classification are greyed out. Unlike statistics, where normally one approach only is chosen, users routinely run several algorithms (like recipes) to see what approach or model provides the best results. For example, for most classification exercises I run a decision tree, Naive Bayes, logistic regression, K-nearest neighbor and support vector machine. Each run only takes one second or less. The results are very useful because it includes: classification accuracy, precision, recall (sensitivity), confusion matrix and area under the curve (AUC). Most healthcare workers are used to seeing results, in terms of sensitivity and specificity. If the results are similar they can be compared in another tab of WEKA called "experimenter" to determine if there is any statistical difference. Furthermore, there is another tab called "select attribute" where the user can determine which attributes really contribute to the outcome, as frequently only a few are really contribute to the outcome. Each algorithm can be further customized to see if the change affects the results. Below is the result of classification to determine what predicts type 2 diabetes in adults. We started with 14 predictors but ended with the best 4. (TP = true positive, FP = false positive) (table 1)Table 1
The "classify" tab is where the fun begins. Selecting the "Choose" option reveals an entire tree of algorithm choices. Those inappropriate for classification are greyed out. Unlike statistics, where normally one approach only is chosen, users routinely run several algorithms (like recipes) to see what approach or model provides the best results. For example, for most classification exercises I run a decision tree, Naive Bayes, logistic regression, K-nearest neighbor and support vector machine. Each run only takes one second or less. The results are very useful because it includes: classification accuracy, precision, recall (sensitivity), confusion matrix and area under the curve (AUC). Most healthcare workers are used to seeing results, in terms of sensitivity and specificity. If the results are similar they can be compared in another tab of WEKA called "experimenter" to determine if there is any statistical difference. Furthermore, there is another tab called "select attribute" where the user can determine which attributes really contribute to the outcome, as frequently only a few are really contribute to the outcome. Each algorithm can be further customized to see if the change affects the results. Below is the result of classification to determine what predicts type 2 diabetes in adults. We started with 14 predictors but ended with the best 4. (TP = true positive, FP = false positive) (table 1)Table 1 For regression, users select the "classify" tab and choose linear regression and for clustering they use the "cluster" tab. Clustering is unsupervised learning where you don't know the classes or attributes so you run clustering to see if there are groups hidden in the data that are worth further exploration. There is also a "select attributes" tab that determines which attributes are most important.I encourage healthcare workers in the data analytics realm to become familiar with WEKA or a similar program, as I believe they will find the programs useful and not that difficult to use.It is true that you must have some background information and most do not want to read an extensive textbook to gain this information. I highly recommend a 2016 ebook (248 page PDF) from Jason Brownlee that teaches basic machine learning and uses WEKA as the software application. It is written simply and clearly and doesn't involve coding or formulas! The cost is only $27 and is well worth it.
For regression, users select the "classify" tab and choose linear regression and for clustering they use the "cluster" tab. Clustering is unsupervised learning where you don't know the classes or attributes so you run clustering to see if there are groups hidden in the data that are worth further exploration. There is also a "select attributes" tab that determines which attributes are most important.I encourage healthcare workers in the data analytics realm to become familiar with WEKA or a similar program, as I believe they will find the programs useful and not that difficult to use.It is true that you must have some background information and most do not want to read an extensive textbook to gain this information. I highly recommend a 2016 ebook (248 page PDF) from Jason Brownlee that teaches basic machine learning and uses WEKA as the software application. It is written simply and clearly and doesn't involve coding or formulas! The cost is only $27 and is well worth it.
Published on September 02, 2016 10:40
No comments have been added yet.



